Clinical Technoshot: in situ Proteomics
Tiziana Bonaldi
Clinical Technoshot Coordinator
| [email protected] | |
| Telephone | 0039 02 94375123 |
| Location | Building 13 Floor 1st Via Adamello 16, Milano |
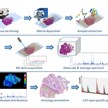
Tumors are characterized by a high degree of heterogeneity that is difficult to access with most analytical methods. The cancer tissue represents the best sample material for any molecular profiling analysis, as it recapitulates all information concerning proteomic and genetic alterations. However, ideally, an analytical method should be able to take advantage of morphological information contained within the cancer tissues, which is not easily possible using standard “shotgun” proteomics approaches. Alternatively, Matrix-Assisted Laser Desorption/Ionization Imaging Mass Spectrometry (MALDI-IMS) is a method that allows the combination of MS analysis with the simultaneous histological evaluation, to profile proteins, peptides and also metabolites in a spatially-resolved fashion. Spatially-resolved MS-measurements of polypeptides (in situproteomics) are directly taken from a tissue section, without destroying it. This combination allows for direct protein analysis of tumor samples, while retaining the spatial architecture and the morphological information contained within the tissues, making it a very valuable tool in cancer research that expand and complement other approaches, such as in situ transcriptomics and metabolomics and Laser Micro Dissection – tandem Mass Spectrometry (LMD-LC-MS/MS)
MALDI-IMS has already proven its potential for various applications in cancer research. Nevertheless, the setup and optimisation of efficient and standardized approaches to make this technique robust and reliable is high challenging.
The goal of this Clinical TechnoShot is to setup and optimize MALDI-IMS approaches at IEO, for the spatially-resolved profiling of proteins and peptides in tumor samples, and integrated in situ proteomic information with similar analysis at the transcriptome level. MALDI-IMS experiment will be also integrated with LMD-LC-MS/MS analyses, that are ongoing and will be also optimized within this Technoshot. The program is therefore focused at developing all the protocols and methods from tissue section up to MALDI-molecular profiles and the in- depth quantitative proteomes achieved from LMD- areas, to assess tissue heterogeneity at the protein level.